Plasmid Profiler Pipeline: Heatmap Display of Plasmid Content in WGS Data
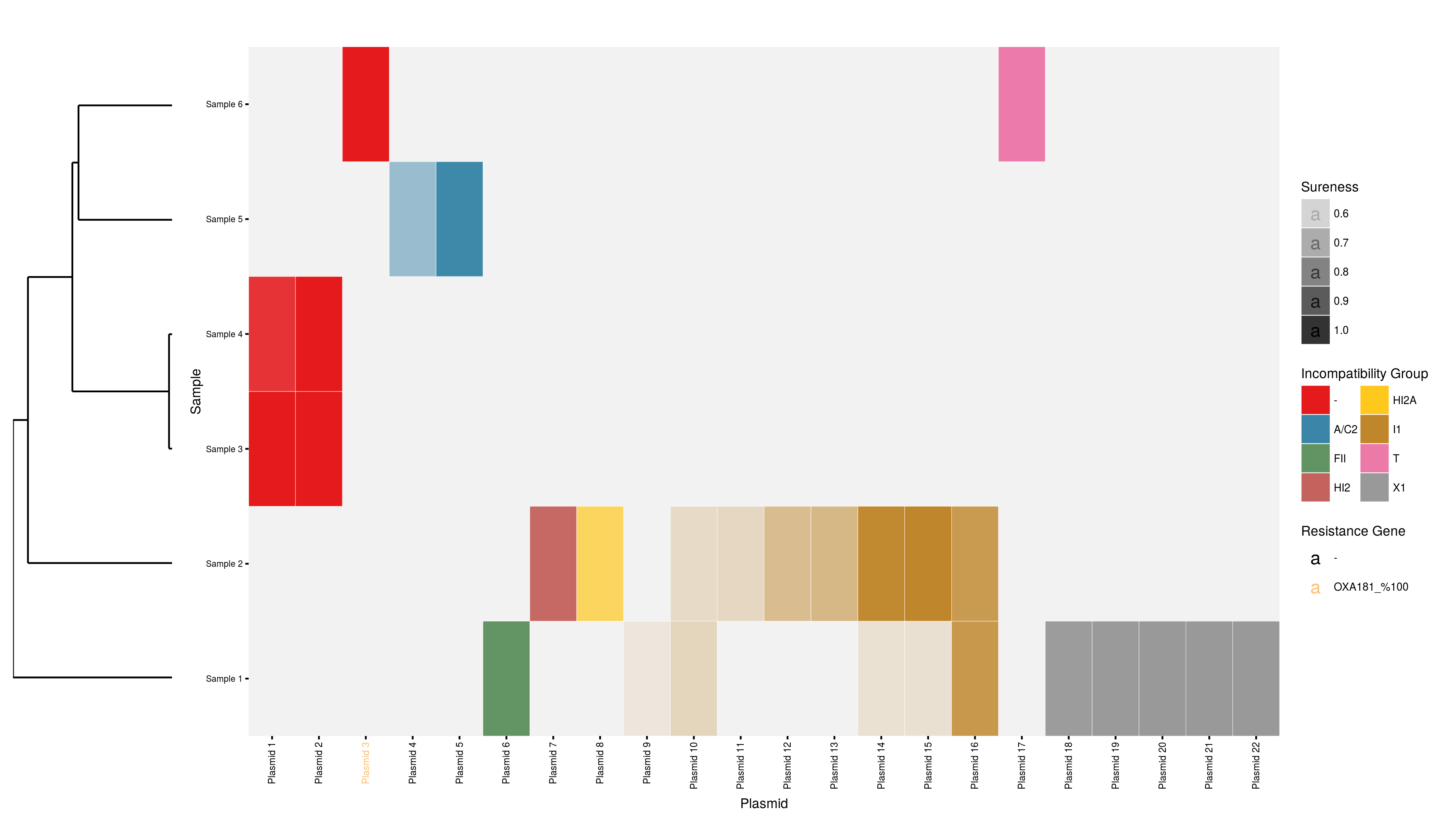
Plasmid profiler is a pipeline to perform comparative plasmid content analysis. It is designed to rapidly bin plasmid content using KAT, Short Read Sequence Typing, and BLAST followed by scoring hits based on a combined measure of maximized coverage and minimized sequence divergence. Hits are then visualized in both static and interactive heatmaps as well as arranged as tabular results. Input is provided in the form of a collection of whole genome sequence reads along with a reference plasmid database and replicon/gene of interest database. The output from the pipeline consists of a png heatmap, an interactive html heatmap, and tabular format results of all plasmids identified and their respective scores.
Operation
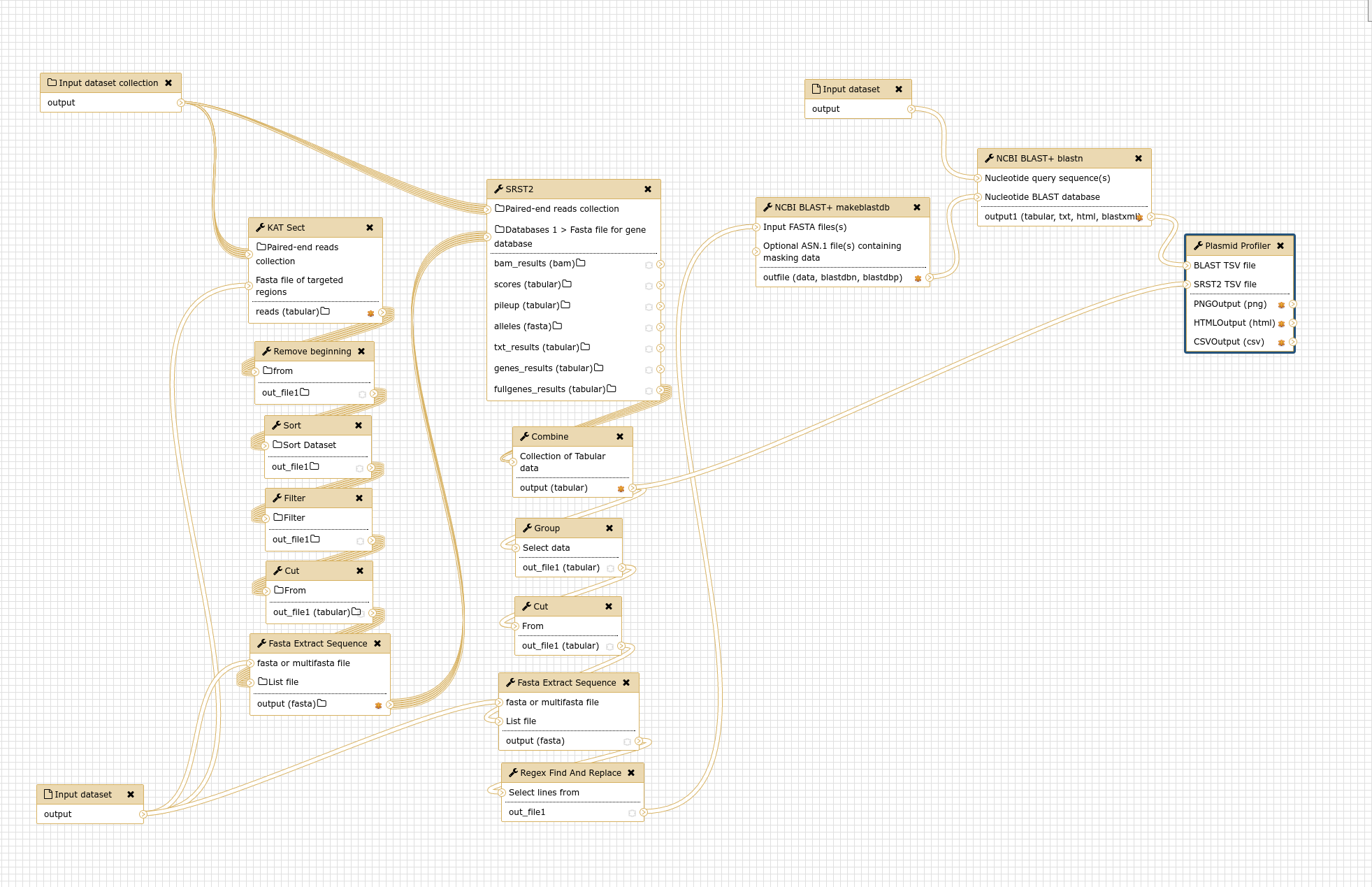
The stages of the Plasmid Profiler pipeline are as follows:
- Preparing input files including:
- A set of sequence reads.
- A reference plasmid database.
- Plasmid Finder replicon database along with genes of interest.
- KAT identifies database plasmids represented in the reads and creates an individualized plasmid database per isolate.
- SRST2 identifies plasmid hits from the individual per isolate databases.
- BLAST identifies the incompatibility groups and genes of interest on hit plasmids.
- Parse and score the identified plasmids using the PlasmidProfiler R package.
- Produce visualizations and export tables using the PlasmidProfiler R package.
Plasmid Profiler is implemented as a Galaxy workflow, with each of these stages implemented using a specific Galaxy tool.
More information on the operation and installation of the pipeline can be found in the Usage and Installation sections.
Contact
Comments, questions, or issues can be sent to:
Adrian Zetner adrian.zetner@canada.ca
Jennifer Cabral jennifer.cabral@canada.ca